GenAge entry for CREBBP (Homo sapiens)
Gene name (HAGRID: 64)
- HGNC symbol
- CREBBP
- Aliases
- RTS; CBP; KAT3A; RSTS
- Common name
- CREB binding protein
Potential relevance to the human ageing process
- Main reason for selection
- Entry selected based on evidence linking the gene product to the regulation or control of genes previously linked to ageing
- Description
A transcriptional regulatory protein, CREBBP acetylates histones and mediates cAMP-gene regulation by binding to phosphorylated CREB1. CREBBP null mice die during embryogenesis, but heterozygous animals exhibit lipodystrophy, have increased insulin (INS) and leptin (LEP) sensitivity, and appear to be protected from weight gain induced by a high-fat diet [429]. In humans, defects in CREBBP have been associated with Rubinstein-Taybi syndrome [428]. Its important roles in transcriptional regulation and its large number of interacting partners suggest CREBBP might potentially be involved in some aspects of ageing.
Cytogenetic information
- Cytogenetic band
- 16p13.3
- Location
- 3,725,055 bp to 3,880,120 bp
- Orientation
- Minus strand
Protein information
- Gene Ontology
-
Process: GO:0000122; negative regulation of transcription from RNA polymerase II promoter
GO:0001666; response to hypoxia
GO:0002223; stimulatory C-type lectin receptor signaling pathway
GO:0006355; regulation of transcription, DNA-templated
GO:0006367; transcription initiation from RNA polymerase II promoter
GO:0006461; protein complex assembly
GO:0006473; protein acetylation
GO:0007165; signal transduction
GO:0007219; Notch signaling pathway
GO:0008589; regulation of smoothened signaling pathway
GO:0016032; viral process
And 15 more GO terms Cellular component: GO:0000123; histone acetyltransferase complex
GO:0000790; nuclear chromatin
GO:0005634; nucleus
GO:0005654; nucleoplasm
GO:0005737; cytoplasm
GO:0016604; nuclear body
Show all GO termsFunction: GO:0000987; core promoter proximal region sequence-specific DNA binding
GO:0001078; transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding
GO:0001085; RNA polymerase II transcription factor binding
GO:0001102; RNA polymerase II activating transcription factor binding
GO:0001105; RNA polymerase II transcription coactivator activity
GO:0001191; transcriptional repressor activity, RNA polymerase II transcription factor binding
GO:0002039; p53 binding
GO:0003682; chromatin binding
GO:0003684; damaged DNA binding
GO:0003700; transcription factor activity, sequence-specific DNA binding
GO:0003713; transcription coactivator activity
And 9 more GO terms
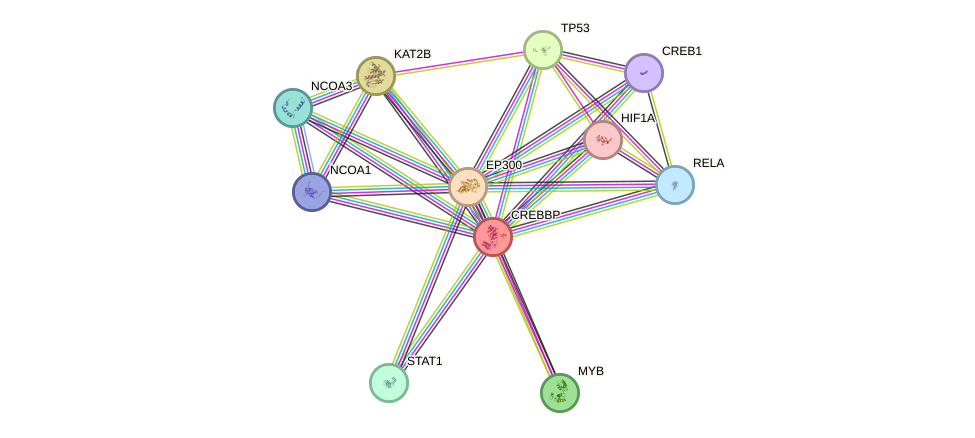
Protein interactions and network
- Protein-protein interacting partners in GenAge
- POU1F1, TP53, WRN, STAT3, AKT1, MYC, FOS, HDAC3, PARP1, BRCA1, CREBBP, HIF1A, NCOR1, EGR1, UBE2I, EP300, PML, HTT, AR, PCNA, XRCC6, FOXO3, FOXO1, HSF1, FOXM1, HDAC1, FGFR1, SP1, JUN, MAPK3, HMGB1, CCNA2, HMGB2, MAP3K5, FOXO4, BMI1, CREB1, ATF2, TBP, HBP1, TFDP1, E2F1, HDAC2, MDM2, DDIT3, RELA, TCF3, SUMO1, HOXB7, NR3C1, CEBPB, ESR1, CTNNB1, ARNTL, CLOCK, PPARGC1A, PPARG, NCOR2, TP73, NFE2L2, CDKN1A, IKBKB
- STRING interaction network
Retrieve sequences for CREBBP
Homologs in model organisms
- Caenorhabditis elegans
- cbp-2
- Caenorhabditis elegans
- cbp-1
- Caenorhabditis elegans
- F40F12.7
- Danio rerio
- crebbpb
- Danio rerio
- crebbpa
- Drosophila melanogaster
- nej
- Mus musculus
- Crebbp
- Rattus norvegicus
- Crebbp
- Saccharomyces cerevisiae
- BDF2
- Saccharomyces cerevisiae
- BDF1
In other databases
- GenAge model organism genes
- A homolog of this gene for Caenorhabditis elegans is present as CELE_F40F12.7
- GenDR gene manipulations
- A homolog of this gene for Caenorhabditis elegans is present as cbp-1